| Python-based Hierarchical ENvironment for Integrated Xtallography |
| Documentation Home |
Generating ligand structures and restraints in the eLBOW GUIOverviewThe electronic Ligand Builder and Optimization Workbench (eLBOW) is the primary tool for generating non-standard ligand restraints in Phenix. In addition to existing as a standalone program, it is also used internally by the LigandFit wizard and phenix.ready_set (integrated with the phenix.refine GUI). In addition to eLBOW, a separate standalone graphical restraint editor is available for advanced customization of restraints and structures. 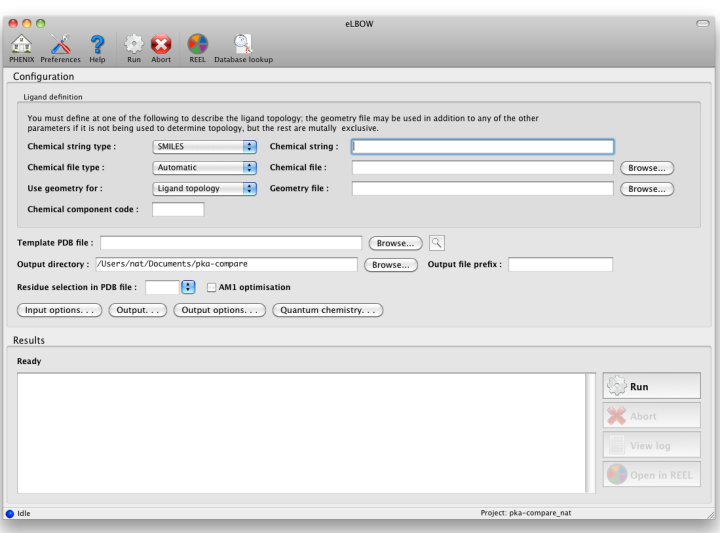 Above: data entry window for eLBOW in the Phenix GUI. Input parameters are grouped separately at top. Input fileseLBOW has extremely flexible options for describing ligand information. The only mandatory data are some description of the molecular topology, i.e. atomic composition and (implicit or explicit) bond connectivity. Four different sources of information may be used:
Only one source of topology information may be provided, but you may also provide the geometry file in addition to one of the first three inputs, to guide eLBOW in constructing the resulting model. You will need to change the drop-down menu labeled "Use geometry for:" to one of the other three options if you do this. The geometry is especially useful if you already have placed the ligand in your structure and now need to refine it. A final source of information is a PDB file containing the template atom names; the geometry and topology information will be discarded. If this is not provided, PHENIX will generate atom names for the output model and CIF file automatically. Output filesBy default, eLBOW will generate a PDB file and a CIF file, suitable for use in any other PHENIX app (or third-party software). Additional output formats are available and may be selected by clicking the "Output. . ." button. You may also manipulate other details such as chirality, ring pucker, and PDB file attributes (including ligand code if you are starting from a SMILES string). 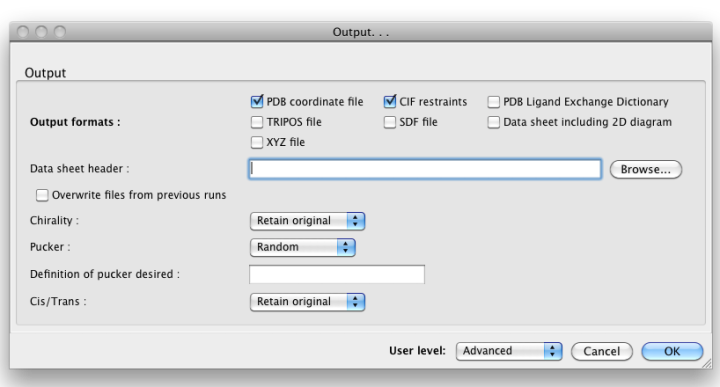 Above: the "Output. . ." dialog. 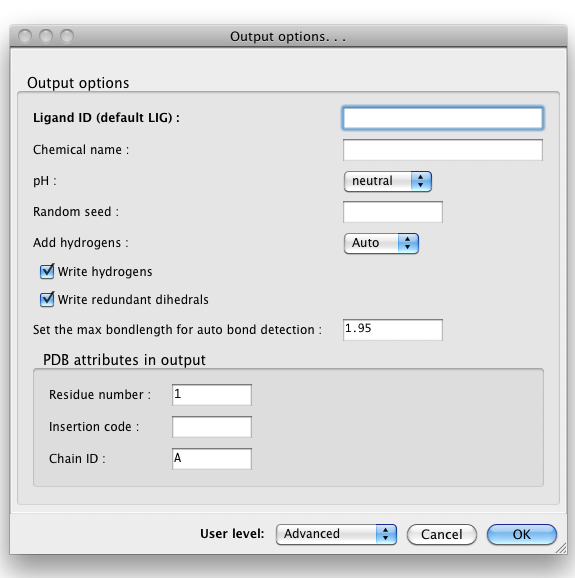 Above: the "Output options. . ." dialog. What to do nextWe recommend opening the output CIF file in REEL to make sure that the results are what you expect. The CIF may be used in any applications requiring restraints as input (primarily phenix.refine, the validation tools, and the AutoBuild and LigandFit wizards). If you have not already placed your ligand in the model, the PDB file is suitable for this, but we recommend trying LigandFit first to save time. References
| |